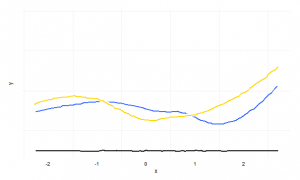
A clever use of geom_smooth(position = ‘jitter’) for plots looking like hand-written.
test.dframe <- data.frame(x = rnorm(100), y = rnorm(100), z = rnorm(100)
theme_xkcd <- theme(
panel.background = element_rect(fill="white"),
axis.ticks = element_line(colour=NA),
panel.grid = element_line(colour="white"),
axis.text.y = element_text(colour=NA),
axis.text.x = element_text(colour="black"),
)
p <- ggplot(data=pleb.clegg, aes(x=Date, y=Pleb))+
geom_smooth(aes(y=Clegg), colour="gold", size=1, position="jitter", fill=NA)+
geom_smooth(colour="white", size=3, position="jitter", fill=NA)+
geom_smooth(colour="dark blue", size=1, position="jitter", fill=NA)+
geom_text(data=pleb.clegg[10, ], family="Humor Sans", aes(x=Date), colour="gold", y=20, label="Searches for clegg")+
geom_text(data=pleb.clegg[22, ], family="Humor Sans", aes(x=Date), colour="dark blue", y=4, label="Searches for pleb")+
geom_line(aes(y=xaxis), position = position_jitter(h = 0.1), colour="black")+
coord_cartesian(ylim=c(-5, 40))+
labs(x="", y="", title="Pleb vs Clegg: Google Keyword Volumes")+
theme_xkcd
http://drunks-and-lampposts.com/2012/10/02/clegg-vs-pleb-an-xkcd-esque-chart/